- Home
- Research
- Optical Molecular Diagnostics and System Technology
Scientific Profile
The Department of Optical Molecular Diagnostics and System Technology is engaged with the development of systems and applications to improve the understanding of infectious diseases. New, innovative approaches, based on optical, molecular and serological multiparameter analyses, will enable faster and more accurate diagnoses as well as detailed epidemiological studies addressing both, the healthcare and the veterinary sectors. In the development of such tests and systems, molecular reference methods such as digital PCR, real-time PCR, next-generation sequencing (NGS) and microarray technologies are used. These techniques are used to evaluate suitable parameters and to optimize them for different applications. Special emphasis is placed on the development of patient-centered point-of-care tests. The entire process of development and translation, including definition of a task, design, research and evaluation, is going to be performed together with industrial partners. Prof. Ralf Ehricht's department focuses on the development and translation of prototypes in the healthcare and veterinary sectors.
Research Topics
- Optical, molecular and serological multiparameter analyses for clinical microbiological diagnostics and epidemiology
- Focus on detection and understanding of antibiotic resistance of pathogenic bacteria
- Epidemiological studies and development of diagnostic tests based on state-of-the-art molecular technologies, such as digital PCR, real-time PCR, Next-Generation-Sequencing (NGS), microarray-based and isothermal analytical methods.
- Microarray-based multiparameter serology using antibody, antigen and peptide arrays.
- Bioinformatics for molecular assay design and analyses of genome sequencing data.
- Building a "bridge" between academic research and industrial partners facilitating the development of innovative diagnostic tests.
- Translation, in cooperation with industrial partners, of research results into commercially available products.
Areas of application

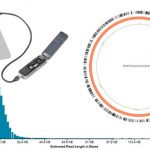
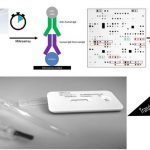
DNA - based multiparameter methods The department of Prof. Ralf Ehricht started its work at Leibniz IPHT in January 2019. Here, as well as previously, its major focus of research was the development and application of various molecular methods for the detection of pathogenic bacteria and their antibiotic resistance properties (MRSA, ESBL, CRE, VRE, etc.) as well as the parallel use of multiple genetic markers for epidemiological purposes, in order to produce genetic fingerprints that could be used for instance in outbreak investigations. In particular, microarray-based methods and specially developed platforms were used (Fig. 1) This included a wide range of technologies and activities including bioinformatic assay design, surface chemistry, manufacturing, test development, image and data evaluation as well as culture-based bacteriological methods under biosafety 2 and 3 conditions. Special attention was given to the implementation of these research results into products that can be used in routine clinical and veterinary settings. Another important topic was to develop methods and procedures for optimal sample preparation and to implement them in complex processes involving real diagnostic samples. Next-Generation-Sequencing - A Tool for the Development of Rapid Tests The MinION platform developed by Oxford Nanopore Technologies (Oxford, UK) is regarded as a next step in next-generation sequencing (Fig. 2). It is based on membrane bound protein nanopores. An ionic current is passed through these pores by setting a voltage across this membrane. If DNA passes through the pore, a change in the current will occur that can be measured and translated into nucleotide sequences. Up to 400 nucleotides per second can be identified per pore, and individual reads can reach lengths of 50,000-100,000 bases. It is possible to quickly and easily analyse bacterial genomes that encompass two or more gigabases. While the length of the generated reads, the comparably low prices and the compact size of the device make the platform attractive, other next-generation sequencing methods (e.g. MySeq, Illumina) still need to be established and applied in parallel in order to proofread nanopore sequences. The Ehricht working group intends to use these technologies to develop new groundbreaking assays on a molecular basis. This is going to be combined with advanced bioinformatics methods as provided by Prof. M. Marz' group. Protein and peptide - based multiparameter methods Serological tests for different pathogens are routinely performed, but as all tests of a panel are usually performed separately and sequentially, costs are rather high and quite a lot of time might pass until the entire panel was processed. In addition, a relatively large amount of sample is required requiring venipuncture. The Ehricht working group was able to show that serological assays can be multiplexed spotting proteins or peptides on microarrays. The IgG levels, and thus the vaccination status, for up to 30 pathogens in parallel can be determined from 1 µl of capillary blood within two hours (Fig. 3). In future, this method could be developed for diagnostic purposes. The antigen panel can quickly be extended or adapted to other applications and resulting assays also can be transferred to point-of-care platforms such as lateral flow (Fig. 3). The method can also be used for antigen/antibody screening or to determine optimal antibody pairings in order to develop and to optimise serological tests.