- Home
- Research
- Photonic Data Science
- Research results
- Machine learning approaches using non-linear multimodal Imaging for biomedical diagnostics
Machine learning approaches using non-linear multimodal Imaging for biomedical diagnostics
18.01.2021
Non-linear multimodal imaging modalities, such as coherent anti-Stokes Raman scattering, two photon excited fluorescence and second harmonic generation, are label-free imaging techniques that provides intrinsic molecular contrast. As these imaging techniques and the resulting data are untargeted, their interpretation requires advanced image analysis methods like machine learning approaches.
By Nairveen Ali // Pranita Pradhan // Elsie Quansah // Hyeonsoo Bae // Tobias Meyer // Michael Schmitt // Jürgen Popp // Thomas W. Bocklitz
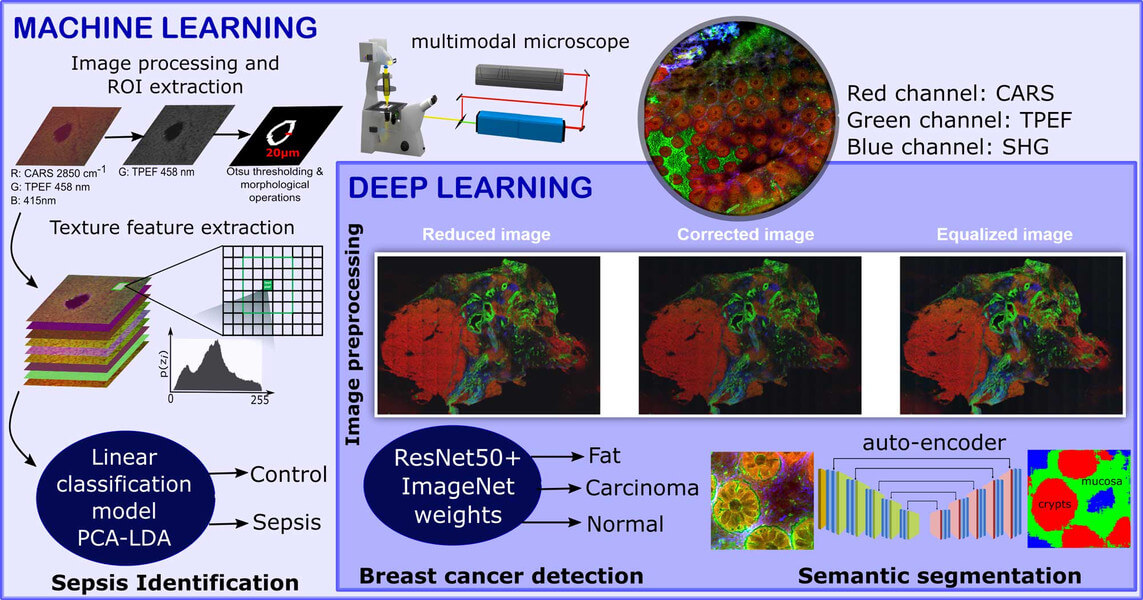
Non-linear multimodal imaging combining coherent anti-Stokes Raman scattering (CARS), two-photon excited autofluorescence (TPEF), and second-harmonic generation (SHG) allows label‐free investigations and biomedical diagnostics. These three individual image modalities provide information about different biomolecules. For example, CARS microscopy can be employed to visualize the distribution of the CH-stretching vibration, TPEF microscopy can measure endogenous fluorophores like NADH, FAD and elastin fibers, and lastly SHG microscopy can provide information about fiber like structures. These non-linear modalities provide a direct visualization of not only the cell and tissue morphology, but also the biomolecular alterations associated to disease progression.
However, these modalities are untargeted which makes the manual interpretation difficult. To circumvent this issue, sophisticated image analysis approaches are required for the interpretation of the acquired image data. In our contributions, the non-linear multimodal images are analyzed using machine learning and deep learning algorithms. We utilize the combination of non-linear multimodal imaging and machine learning/deep learning to solve different biomedical tasks including the detection of breast cancer (Ali, 2019), the early identification of sepsis (Yarbakht, 2019) and the semantic segmentation of crypt structures for a characterization of inflammatory bowel diseases (Bocklitz, 2019).
The identification of sepsis in a mouse model of peritonitis was carried out using a classical machine learning model that combined extracted texture features of the non-linear multimodal images, a principal component analysis and a linear discriminant analysis (PCA-LDA). In contrast, an automatic detection of breast cancer was achieved using a deep learning approach. This approach used a fully connected convolutional neural network called ResNet50. Here, the pretrained version of the ResNet50 network on the ImageNet dataset was fine-tuned to identify the breast cancer regions via patch-based convolutional neural network approach. Although the sample sizes of the above-mentioned tasks were small, machine learning algorithms showed promising identification results for sepsis and breast cancer. The performance of the two image classification tasks was evaluated using the class sensitivity or the area under the curve (AUC) of a receiver operating characteristics curve.

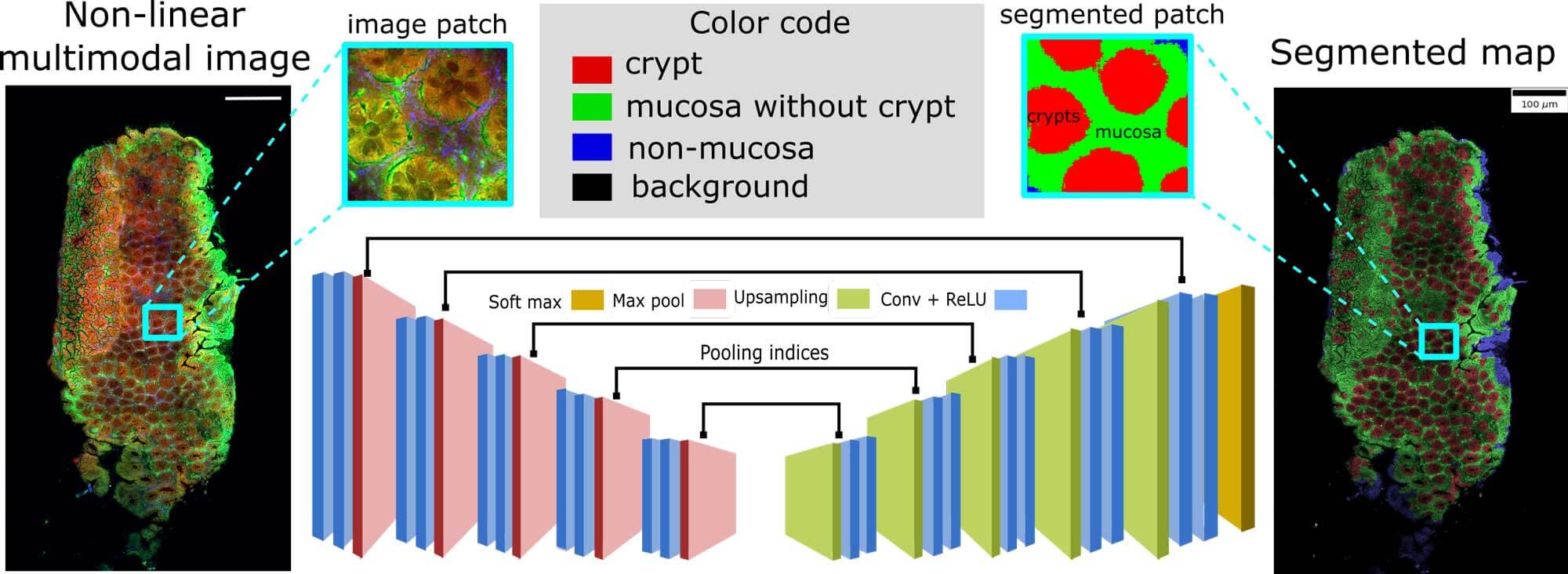
Semantic segmentation of crypts in non-linear multimodal images was performed by an advanced auto-encoder model (SegNet). The input to the SegNet model was a non-linear multimodal image and the output was a segmented image with four different regions: background, non-mucosa, mucosa without crypts and crypts. These regions especially the mucosa region and the crypts are crucial indicators for inflammatory bowel disease (IBD). Its automatic segmentation without any manual intervention can lead to automatic characterization of IBD.
Overall, the non-invasive nature of this imaging technique enables in vivo measurements, which offers a low‐risk diagnostic approach for various diseased. If adequate machine learning / deep learning approaches are combined with these non-linear multimodal imaging techniques, an automatic decision-making system for biomedical diagnostics including the detection of breast cancer (Ali, 2019), the early identification of sepsis (Yarbakht, 2019) and the semantic segmentation of crypt structures for a characterization of inflammatory bowel diseases (Bocklitz, 2019) can be achieved.
